Caixia Gao Team Achieves Megabase-Scale Precision Genome Editing in Eukaryotic Cells
Prof. Caixia Gao's team from the Institute of Genetics and Developmental Biology of the Chinese Academy of Sciences has developed two advanced genome editing platforms,collectively known as Programmable Chromosome Engineering (PCE) systems,enabling precise and megabase-scale manipulation of plant chromosomes.
Our study,published online in Cell on August 4 (https://doi.org/10.1016/j.cell.2025.07.011),illustrates that PCE systems enable diverse,high-precision DNA manipulations—from kilobase- to megabase-scale—in higher organisms,especially plants.
Extensive research has demonstrated the immense potential of the site-specific recombinase Cre-Lox system for precise chromosomal manipulation. However,its broader application has been hindered by three critical limitations: (1) reversible recombination reactions—stemming from the inherent symmetry of Lox sites—can negate desired edits;(2) the tetrameric nature of Cre recombinase complicates engineering efforts,hindering activity optimization;and (3) residual Lox sites after recombination may compromise editing precision.
Our research team addressed each of these challenges and developed novel methods to advance the state of this technology. To begin with,we built a high-throughput platform for rapid recombination site modification and proposed an asymmetric Lox site design. This approach developed novel Lox variants that reduce reversible recombination activity by over 10-fold (approaching the background level of negative controls) while retaining high-efficiency forward recombination.
We then leveraged their recently developed AiCE (AI-informed Constraints for protein Engineering) model—a protein-directed evolution system integrating general inverse folding models with structural and evolutionary constraints—to develop AiCErec,a recombinase engineering method. This approach enabled precise optimization of Cre’s multimerization interface,yielding an engineered variant with a recombination efficiency 3.5 times that of wild-type Cre.
Furthermore,we designed and refined a scarless editing strategy for recombinases. By harnessing the high editing efficiency of prime editors,we developed Re-pegRNA,a method that uses specifically designed pegRNAs to perform re-prime editing on residual Lox sites,precisely replacing them with the original genomic sequence,thereby ensuring seamless genome modifications.
The integration of these three innovations led to the creation of two programmable platforms,PCE and RePCE. These platforms allow flexible programming of insertion positions and orientations for different Lox sites,enabling precise,scarless manipulation of DNA fragments ranging from kilobase to megabase scale in both plant and animal cells. Key achievements include: targeted integration of large DNA fragments up to 18.8 kb,complete replacement of 5-kb DNA sequences,chromosomal inversions spanning 12 Mb,chromosomal deletions of 4 Mb,and whole-chromosome translocations.
As a proof of concept,we used this technology to create herbicide-resistant rice germplasm with a 315-kb precise inversion,showcasing its transformative potential for genetic engineering and crop improvement.
Our pioneering work not only overcomes the historical limitations of the Cre-Lox system but also opens new avenues for precise genome engineering in a variety of organisms.
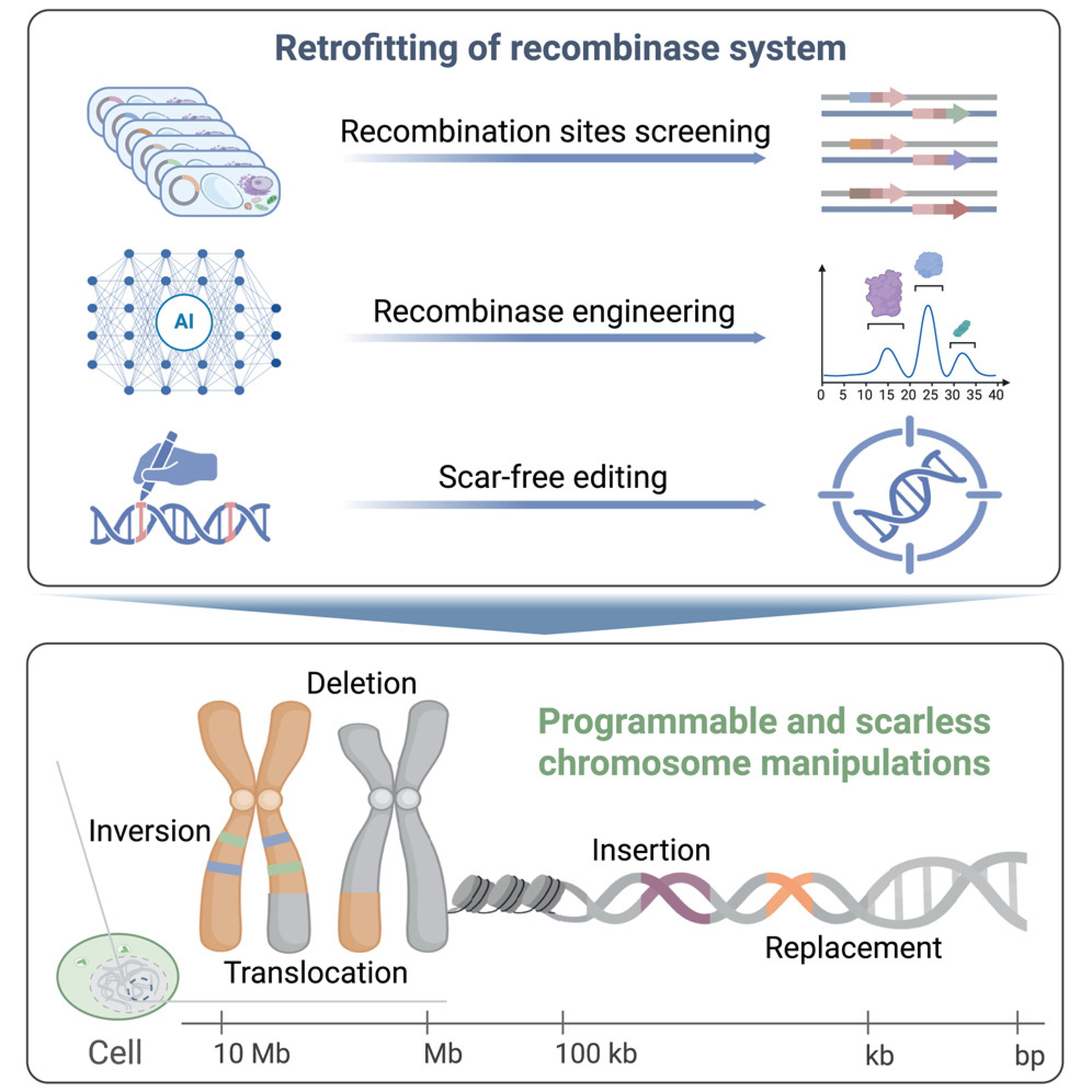
Figure: A description of the development of PCE systems and accurate chromosome editing.
File Download:
